CBS News Live
CBS News Texas: Local News, Weather & More
Watch CBS News





Scattered storms could continue into Friday morning.

Five North Texas nonprofits are competing for more than $200,000 in funding from the United Way of Metropolitan Dallas.

For some it's sport, for others, it's science: chasing storms. And for everyone who loves the rush, it's the prime time of the year to head into danger.

A criminal investigation is unfolding in Denton after a 13-year-old girl is attacked on a school bus by adults.

Neighbors in the Las Vegas Trail neighborhood said they're shaken up and worried about everyone's safety following the shooting Wednesday night.

A Dallas police source said the sexual assault investigation is over due to insufficient evidence.

There is no question that Nehls served overseas and engaged in combat, but military documents show he received one Bronze Star instead of two.

A North Texas mother partnered with the Texas Health Resources Foundation to provide a potentially life-saving tool to at-risk pregnant and postpartum moms.

A jury sentenced Jerry Don Elders to death for the murder of a woman he kidnapped while on the run.

Branson ran over a mile in the dark, through downed power lines and debris to get help.

Tickets are not officially on sale yet. You can sign up for presale tickets on the AT&T Stadium website.

Sadly, a truck driver was killed when his 18-wheeler overturned.
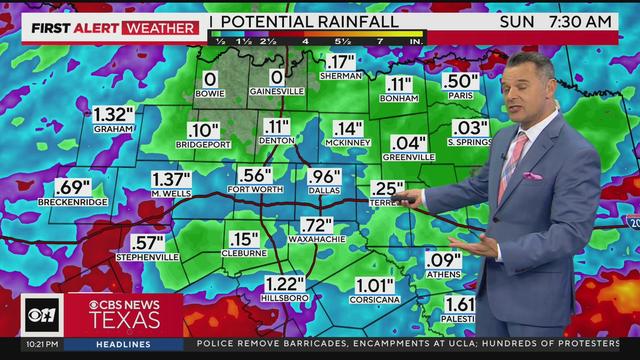
The flood watch was allowed to expire, but with some more widespread rain/storm chances Saturday into Sunday it's possible that another watch will be issued at some point this weekend.

For some it's sport, for others, it's science: chasing storms. And for everyone who loves the rush, it's the prime time of the year to head into danger.

Five North Texas nonprofits are competing for more than $200,000 in funding from the United Way of Metropolitan Dallas.

WARNING: Content may be disturbing for some. A criminal investigation is unfolding in Denton after a 13-year-old girl is attacked on a school bus by adults.

It turns out more water wasters are turning up in Texas. Our media partners at Texas Monthly show us how the trend is now rising across the state.

Scattered storms could continue into Friday morning.

The flood watch was allowed to expire, but with some more widespread rain/storm chances Saturday into Sunday it's possible that another watch will be issued at some point this weekend.

On Friday, a few early morning showers and isolated storms are possible.
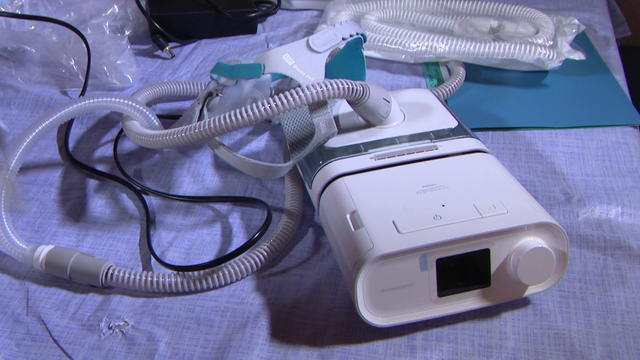
In a statement, Philips said it does not admit any wrongdoing but chose to settle "to end the uncertainty associated with litigation in the U.S."

A scammer a North Texas woman met on Instagram claimed to be a German cardiologist, and for months, the two messaged back and forth, building what she thought was a true relationship.

Texas police departments have the discretion to determine the frequency and extent of additional driving training for their officers. While some require driving training yearly or every other year, others do not.

Some departments opt to melt the firearms down, while others choose to crush them. However, there are instances where firearms, or at least parts of them, escape destruction altogether.

Several police departments told the CBS News Texas I-Team they were unaware of this practice, even though it was stated in the contracts they signed with the company, Gulf Coast GunBusters.

Luka Doncic scored 20 of his 35 points in the second half and added 10 assists and seven rebounds, propelling the Dallas Mavericks to a 123-93 victory over the Los Angeles Clippers in Game 5 and a 3-2 lead in the first-round series.

Jason Robertson scored a power-play goal to put Dallas ahead late in the second period after Tyler Seguin took a shot to the face and the Stars beat the Vegas Golden Knights 3-2 in Game 5.

The running back is officially a Dallas Cowboy again after a steak dinner that sealed the deal.

Tickets are not officially on sale yet. You can sign up for presale tickets on the AT&T Stadium website.

Wyatt Johnston scored his third goal in two games, and the Dallas Stars beat the defending champion Vegas Golden Knights 4-2 to even their first-round NHL playoffs series.

Scattered storms could continue into Friday morning.

Five North Texas nonprofits are competing for more than $200,000 in funding from the United Way of Metropolitan Dallas.

For some it's sport, for others, it's science: chasing storms. And for everyone who loves the rush, it's the prime time of the year to head into danger.

A criminal investigation is unfolding in Denton after a 13-year-old girl is attacked on a school bus by adults.

Neighbors in the Las Vegas Trail neighborhood said they're shaken up and worried about everyone's safety following the shooting Wednesday night.

In a statement, Philips said it does not admit any wrongdoing but chose to settle "to end the uncertainty associated with litigation in the U.S."

A scammer a North Texas woman met on Instagram claimed to be a German cardiologist, and for months, the two messaged back and forth, building what she thought was a true relationship.

They found him guilty – now four jurors are explaining how they were convinced to convict Dr. Raynaldo Ortiz.

Texas police departments have the discretion to determine the frequency and extent of additional driving training for their officers. While some require driving training yearly or every other year, others do not.

Some departments opt to melt the firearms down, while others choose to crush them. However, there are instances where firearms, or at least parts of them, escape destruction altogether.

There is no question that Nehls served overseas and engaged in combat, but military documents show he received one Bronze Star instead of two.

President Biden said "no," the National Guard should not intervene in the protests.

Keith Davidson, a Los Angeles-based lawyer, told jurors about how he represented Stormy Daniels in talks with Michael Cohen.

The Biden administration said it's erasing debt for people who attended the for-profit Art Institutes, which shut down in September.

The Biden administration is considering bringing certain Palestinians fleeing war-torn Gaza to the U.S. as refugees, according to internal federal government documents obtained by CBS News.

Self-driving 18-wheelers have longtime truckers worried about their livelihood and others concerned that the technology needs more testing to make sure the public is safe.

McDonald's concept restaurant CosMc's has taken its drink-focused menu to Dallas for its second-ever location.

With the country on the cusp of greeting the return of spring, a warm-weather treat is once again available for free for a limited time only.

Kelli and Michael Regan were looking for a new dog. The breeder they found online asked them to pay with gift cards.

Target, looking for ways to add sales, is relaunching its Target Circle loyalty program including a new paid membership with unlimited free same-day delivery in as little as an hour for orders over $35.

A North Texas mother partnered with the Texas Health Resources Foundation to provide a potentially life-saving tool to at-risk pregnant and postpartum moms.

Plaintiffs have three months to vote on whether to approve a proposed legal settlement that would resolve nearly all talc lawsuits.

Cat deaths and neurological disease are "widely reported" around farms where the H5N1 bird flu virus was detected, health officials say.

The White House had been due to decide on the menthol cigarette rule in March.
First known HIV cases from a nonsterile injection for cosmetic reasons highlights the risk of unlicensed providers.

Within hours of the vote, the U.S. Chamber of Commerce announced it would sue to block the ban. Dallas employment attorney Rogge Dunn predicts employees will ultimately win this battle.

The closure affects both Dom's locations in Chicago, and all 33 Foxtrot stores in Chicago, Texas, and the Washington D.C. area.

Texas law SB 14 prohibits drug and surgical "gender transition" interventions for minors.

The projects are expected to create at least 17,000 construction jobs and 4,500 manufacturing jobs.

After more than 40 years in business, 99 Cents Only Stores, a discount chain, announced on Thursday that it will close all 371 of its locations and cease operations.

Luka Doncic scored 20 of his 35 points in the second half and added 10 assists and seven rebounds, propelling the Dallas Mavericks to a 123-93 victory over the Los Angeles Clippers in Game 5 and a 3-2 lead in the first-round series.

Jason Robertson scored a power-play goal to put Dallas ahead late in the second period after Tyler Seguin took a shot to the face and the Stars beat the Vegas Golden Knights 3-2 in Game 5.

The running back is officially a Dallas Cowboy again after a steak dinner that sealed the deal.

Tickets are not officially on sale yet. You can sign up for presale tickets on the AT&T Stadium website.

Wyatt Johnston scored his third goal in two games, and the Dallas Stars beat the defending champion Vegas Golden Knights 4-2 to even their first-round NHL playoffs series.

Tickets are not officially on sale yet. You can sign up for presale tickets on the AT&T Stadium website.

Paramount said long-time CEO Bob Bakish will leave the company, which is in discussions to explore a sale or merger.

The vinyl sales alone were monumental, Billboard said, with "the largest sales week for an album on vinyl in the modern era."

Harvey Weinstein's 2020 conviction on felony sex crime charges has been overturned by the State of New York Court of Appeals.

Taylor Swift fans have found a way to feel "a little bit closer to" their hero at a London watering hole, and The Black Dog pub is lapping it up.

The flood watch was allowed to expire, but with some more widespread rain/storm chances Saturday into Sunday it's possible that another watch will be issued at some point this weekend.

For some it's sport, for others, it's science: chasing storms. And for everyone who loves the rush, it's the prime time of the year to head into danger.

Five North Texas nonprofits are competing for more than $200,000 in funding from the United Way of Metropolitan Dallas.

WARNING: Content may be disturbing for some. A criminal investigation is unfolding in Denton after a 13-year-old girl is attacked on a school bus by adults.

It turns out more water wasters are turning up in Texas. Our media partners at Texas Monthly show us how the trend is now rising across the state.

Dallas artist Roberto Marquez traveled to the Rafah Crossing in Egypt, the U.S. capital and will attend this weekend's statewide protest in Austin.

On Friday, hundreds of thousands of fans gathered outside and all around Globe Life Field in Arlington to celebrate the Texas Rangers historical World Series win!

Babies in the neonatal intensive care unit at several Texas Health hospitals were dressed in creative costumes for Halloween.

Is that the smell of cotton candy, beignets and brisket wafting over Fair Park? It sure is, and we are here for it!

No one puts these dolls back in their boxes. Babies in the neonatal intensive care unit at Texas Health Harris Methodist Hospital Southwest Fort Worth are pretty in pink!